The «Epigenetic Regulations and Seed Development» team focuses on mechanisms underlying the fine-tuned orchestration of developmental transitions during plant female reproductive development.
We explore the regulation and the functional role of heterochromatin, and specification of cellular identities. Our model systems include Arabidopsis, maize and apomictic Paspalum. We combine modern technologies for nucleic acids sequencing, biomolecule delivery, and imaging of live tissues to understand developmental plasticity and transgenerational inheritance in plants. Besides fundamental interest, this work could contribute to advancing innovations in plant biotechnologies, such as asexual reproduction, haploidization, manipulation of recombination events, and novel epiallelic variation.
Establishment and dynamics of epigenetic marks:
Establishing distinct epigenome landscapes and gene expression patterns is essential for reproductive success and genome inheritance. We study DNA (hemi)methylation dynamics throughout reproductive development in Arabidopsis and maize by using epigenomics and the visualization tools we have recently designed and validated (Ingouff et al., 2017). The approaches contribute to better understand inheritance of DNA methylation patterns and to define the modalities of DNA reprogramming occurring in plant reproductive cell types. In particular, interactions with cell division machinery and the well-known, yet poorly understood, phenomenon of paramutation in maize are explored.
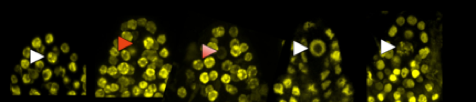
Picture legend:Chromatin dynamics within young ovules of Arabidopsis. The arrows indicate the variations (white + / red -) observed in the nucleus of the female germline. (Picture: M. Ingouff).
Genomics and epigenomics of apomixis:
Although a central role of epigenetic alterations has been hypothesized for its emergence, our understanding of the molecular and cellular modulations underlying apomictic developments remains scarce. Previous works revealed that apomictic Paspalum genomes harbor a specific, hemizygous, highly heterochromatic region of unknown origin and functional role. Using transcriptional resources already available and current efforts to generate genomic assemblies of sexual and apomictic individuals, our objective is to determine its nature and resolve its origin and function.
Cellular identities, plasticity and canalization of reproductive cell fate:
Using genetic and cell biology approaches, we characterize mutations affecting cellular identities during female reproduction in Arabidopsis or maize. In particular, we focus on members of the RdDM pathway and on homologs of genes identified in Paspalum apomicts and encoding factors involved in noncoding RNA methylation. In addition, using imaging and modeling approaches, we address the functional link between ovule architecture and the plasticity and canalization of female germ cell specification in early ovules.
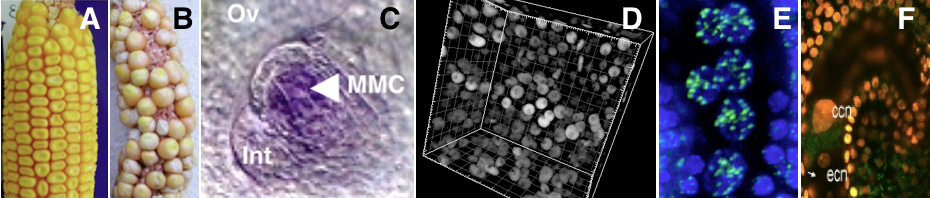
Picture legend: A-B. maize,sterility phenotype of ago104 mutant. C. maize : expression pattern of Dmt102 DNA methyltransferase. D. Arabidopsis : histone marker in early ovule. E. Arabidopsis: histone methylation mark during gametogenesis. F.Arabidopsis : histone turn over in mature gametophyte. (Pictures: D.Grimanelli; M.Garcia-Aguilar; D.Autran; M.Pillot; M.Ingouff).
Partnerships:
The team collaborates on international projects with overseas partners from Mexico, (JEAI EpiMaize U. of Veracruz ; LANGEBIO/CINVESTAV, Irapuato), Argentina (U. Nacional de Rosario, Rosario ; U. Nacional del Sur, Bahia Blanca; IBONE, Corrientes), USA (Cold Spring Harbor Laboratory), and Europe (U. Hamburg, U. Zürich, U. Milano, et U. Perugia). We are also involved in initiatives and projects with national laboratories (IBMP, IBP, RDP/ENS Lyon, CNRS-Paris 7, AGAP). Recently, this network was consolidated by two European Marie Skłodowska-Curie actions, two Agropolis Foundation grants for international mobility, and two ANR international collaborative research projects. .
Education:
We participate in education by training Master and PhD students, participating in plant sciences Master programs at the University of Montpellier and other institutions (University of Orsay XI, ENS Paris), and coordinating the Plant Epigenetics course as part of the Plant Functional Biology Master program at the University of Montpellier.
On-going projects:
H2020 MSCA-RISE MAD - Mechanisms of Apomictic Development
ANR-DFG CHROMOBREED - Elucidating the role and regulation of heterochromatin during maize reproduction, and exploring its potential for developing novel plant breeding strategies
Agropolis Foundation APOMOVIE - Live Imaging of Reproductive Development in Sexual and Apomictic Grasses
Agropolis Fondation EPIMAIZE - Innovative Tools to Study Trans-generational Epigenetic Inheritance in Maize
Agropolis Foundation RNAPO - RNA methylation and reproductive modes in plants: characterization of molecular factors involved in the transition between sexual reproduction and apomixis
ANR REMET - Dynamics of Eukaryotic DNA Remethylation
H2020 MSCA-FELLOWSHIPS EPIMAIZE - Understanding the Maize Epigenome and its Role in Development
H2020 MSCA-RISE PROCROP - Harnessing Plant Reproduction for Crop Improvement
ANR-SNF IMAGO - Imaging and Modelling Growth of Plant Ovules



